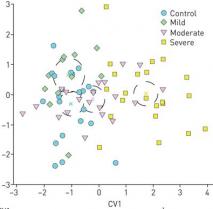
Our research involves applying mass spectrometry-based technologies to perform molecular phenotyping at the population level. This work entails the development of both LC/MS and GC/MS methods for the quantification of lipids and other metabolites. A major focus of the lab also centers on developing the associated bioinformatics approaches including pathway analysis and network construction as well as multivariate statistical methods for integration of 'omics-based data structures.
Lab news
KI News
PubMed
- Recent Developments in Single-Cell Metabolomics by Mass SpectrometryA Perspective
- Mechanistic in vitro study of the effect of Cucurbita moschata (Cucurbitaceae) on carbohydrate digestive enzymes
- The Application of Untargeted Metabolomic Approaches for the Search of Common Bioavailable Metabolites in Human Plasma Samples from Lippia citriodora and Olea europaea Extracts
- Modification of xylan in secondary walls alters cell wall biosynthesis and wood formation programs and improves saccharification
- Attenuated kidney oxidative metabolism in young adults with type 1 diabetes
VR News
- This RSS feed has been discontinued. Welcome to our website Vetenskapsradet.se where you can subscribe to updates and newsletters.
- This RSS feed has been discontinued. Welcome to our website Vetenskapsradet.se where you can subscribe to updates and newsletters.
- MATRAC 1 School - time to apply
- Decision within Conference grants
- Open call - JPIAMR Network Call on Virtual Research Institute
Metabolomics News
- METASPACE-ML: Context-specific metabolite annotation for imaging mass spectrometry using machine learning - Nature.com
- Severe anemia in preterm infants associated with increased bacterial virulence potential and metabolic disequilibrium - Nature.com
- The Human Protein Atlas Launches Version 24 - Technology Networks
- Comorbidities confound metabolomics studies of human disease - Nature.com
- Gut Microbiota and Metabolomics Unveil the Mechanisms of Lomatogonium Rotatum in Ameliorating Visceral Fat and Serum Lipids in High-Fat Diet-Induced Obese Mice - Frontiers